There are many sources of water quality monitoring data including instruments (ex: YSI instruments) and open source data sets (ex: USGS and NDBC), all of which are susceptible to errors/inaccuracies due to drift. driftR provides a grammar for cleaning and correcting these data in a “tidy”, reproducible manner.
What’s New
Version 1.1 of driftR is here! It includes:
- a streamlined
dr_readfunction that includes built-in support for YSI Sonde 6600, YSI EXO, and Onset HOBO products. - new aregument in
dr_readthat gives the option to read in only clean variable names (e.g., no special characters, no spaces, etc.) using functionality from thejanitorpackage. - expanded functionality for
dr_drop, including the ability to drop by date range and by using expressions. - ability to convert observations that are likely measurement errors to
NAusing either a date range or by using an expression using a new function calleddr_replace. - changes under the hood to
dr_factorthat expand it ability to handle a variety of date formats automatically.
Installation
The easiest way to get driftR is to install it from CRAN:
You can also install the development version of driftR from Github with devtools:
Background
The driftR package implements a series of equations used in Dr. Elizabeth Hasenmueller’s hydrology and geochemistry research. These equations correct continuous water quality monitoring data for incremental drift that occurs over time after calibration. There are two forms of corrections included in the package - a one-point calibration and a two-point calibration. One-point and two-point calibration values are suited for different types of measurements. The package is currently written for the easiest use with YSI multiparameter Sonde V2 series products, YSI EXO products, and Onset HOBO products.
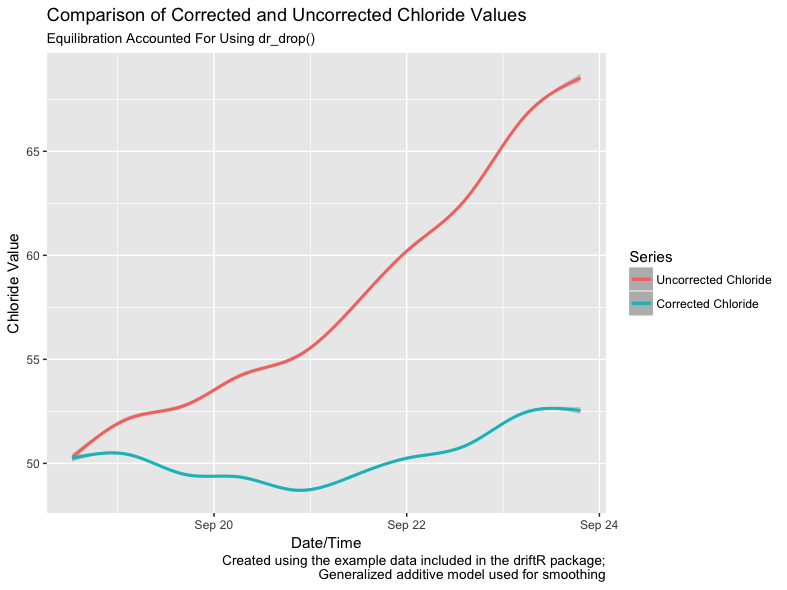
The figure below illustrates the difference in chloride values between the uncorrected data and the same data with the drift corrections implemented by driftR applied. Note that the uncorrected data drifts to higher values over time. driftR uses calibration data to correct this drift.
Usage
As shown, continuous water quality instruments drift over time, so it becomes necessary to correct the data to maintain accuracy. driftR provides five verbs for applying these corrections in a consistent, reproducible manner: read, factor, correct, drop, and replace. These verbs are designed to be implemented in that order, though there may be multiple applications of correct for a given data set. All of the core functions for driftR have the dr_ prefix, making it easy to use them interactively in RStudio.
Basic Use
The following example shows a simple workflow for applying these verbs to some hypothetical data:
# load the driftR package
library(driftR)
# import data exported from a Sonde
waterTibble <- dr_read(file = "data.csv", instrument = "Sonde", defineVar = TRUE,
cleanVar = TRUE, case = "snake")
# calculate correction factor and keep dateTime var
# results stored in new vector corrFac and dateTime
waterTibble <- dr_factor(waterTibble, corrFactor = corrFac, dateVar = Date,
timeVar = Time, keepDateTime = FALSE)
# apply one-point calibration to SpCond;
# results stored in new vector SpConde_Corr
waterTibble <- dr_correctOne(waterTibble, sourceVar = SpCond, cleanVar = SpCond_Corr,
calVal = 1.07, calStd = 1, factorVar = corrFac)
# apply two-point calibration to pH;
# results stored in new vector pH_Corr
waterTibble <- dr_correctTwo(waterTibble, sourceVar = pH, cleanVar = pH_Corr,
calValLow = 7.01, calStdLow = 7, calValHigh = 11.8,
calStdHigh = 10, factorVar = corrFac)
# drop observations to account for instrument equilibration
waterTibble <- dr_drop(waterTibble, head=10, tail=5)
#replace observations with NA for a date range
waterTibble <- dr_replace(waterTibble, sourceVar = pH, overwite = TRUE, dateVar = Date,
timeVar = Time, from = "2018-02-05", to = "2018-02-09")
Use with %>%
All of the core functions return tibbles (or data frames) and make use of the tidy evaluation pronoun .data, so using them in concert with the pipe (%>%) is straightforward:
# load the driftR package
library(driftR)
# import data exported from a Sonde
waterTibble <- dr_read(file = "sondeData.csv", instrument = "Sonde", defineVar = TRUE,
cleanVar = TRUE, case = "snake")
# caclulate correction factors, apply corrections, drop observations, and replace observations
waterTibble <- waterTibble %>%
dr_factor(corrFactor = corrFac, dateVar = Date, timeVar = Time,
keepDateTime = TRUE) %>%
dr_correctOne(sourceVar = SpCond, cleanVar = SpCond_Corr, calVal = 1.07,
calStd = 1, factorVar = corrFac) %>%
dr_correctTwo(sourceVar = pH, cleanVar = pH_Corr, calValLow = 7.01, calStdLow = 7,
calValHigh = 11.8, calStdHigh = 10, factorVar = corrFac) %>%
dr_drop(head=10, tail=5) %>%
dr_replace(waterTibble, sourceVar = pH, overwite = TRUE, dateVar = Date,
timeVar = Time, from = "2018-02-05", to = "2018-02-09")Additional Documentation
See the package website for more information on these functions and a detailed vignette describing how to get started with driftR. There is also an additional vignette describing the specific ways in which dates and times can be used in driftR functions.
We also provide some introductory examples for how to use tidyr, ggplot2, and several other R packages to conduct some initial exploratory data analysis of driftR output. Finally, we provide a third vignette designed for users of instruments not supported directly by driftR who wish to use driftR with their data.
You can also view the help files from within R:
Want to Contribute?
Have a Concern?
If driftR does not seem to be working as advertised, please help us creating a reproducible example, or reprex, that makes it easy to get help. You can find additional details in our support document.
Adding dr_read Functionality
We are interested in expanding the built-in capabilities of driftR to read in water quality data from other sources. As of version 1.1, we provide built-in support for YSI Sonde 6600, YSI EXO, and Onset HOBO products.
If you have some sample data (~500 observations are ideal) from another model or brand of instrument and are willing to share it, please reach out to one of the package authors or, better yet, open an Issue. If you have some R skills and want to write the function yourself, feel free to check out our contributing document and fork driftR.
Contributor Code of Conduct
Please note that this project is released with a Contributor Code of Conduct. By participating in this project you agree to abide by its terms.

